|
SpliceDB is available at the computational genomic Web server of the Sanger Centre. Exploiting two different genes for evaluation of metrics of. We also built weight matrices for the major splice groups, which can be incorporated into gene prediction programs. Moreover, we decided to use as our negative dataset, 64 experimentally validated intronic BRCA variants, all of which are located outside the consensus splice site, in nucleotide positions ranging from 13 to 113 at the 3 splice site and from +9 to +104 at the 5 splice site. We have carried out an initial analysis of the dynamics of the recent evolution of the splice-sites sequences on a large collection of human, rodent (mouse. The information about verified splice site sequences for canonical and non-canonical sites is presented in SpliceDB with the supporting evidence. They can be classified after sequence analysis as: 79 GC-AG pairs (of which one was an error that corrected to GC-AG), 61 errors corrected to GT-AG canonical pairs, six AT-AC pairs (of which two were errors corrected to AT-AC), one case was produced from a non-existent intron, seven cases were found in HTG that were deposited to GenBank and finally there were only two other cases left of supported non-canonical splice pairs. Most 5' splice sites start with GTRAG, and remaining ones have a stronger preference for AG at the end. Out of 171 human non-canonical and EST-supported splice pairs, 156 (91.23%) had a clear match in the human HTG. than 99 of human introns are spliced by the major U2-dependent spliceosome. To check the conservative dinucleotides we compared sequences of human non-canonical splice sites with sequences from the high throughput genome sequencing project (HTG). EST alignments allowed us to verify only the exonic part of splice sites. We especially studied non-canonical splice sites, which comprise 3.73% of GenBank annotated splice pairs. 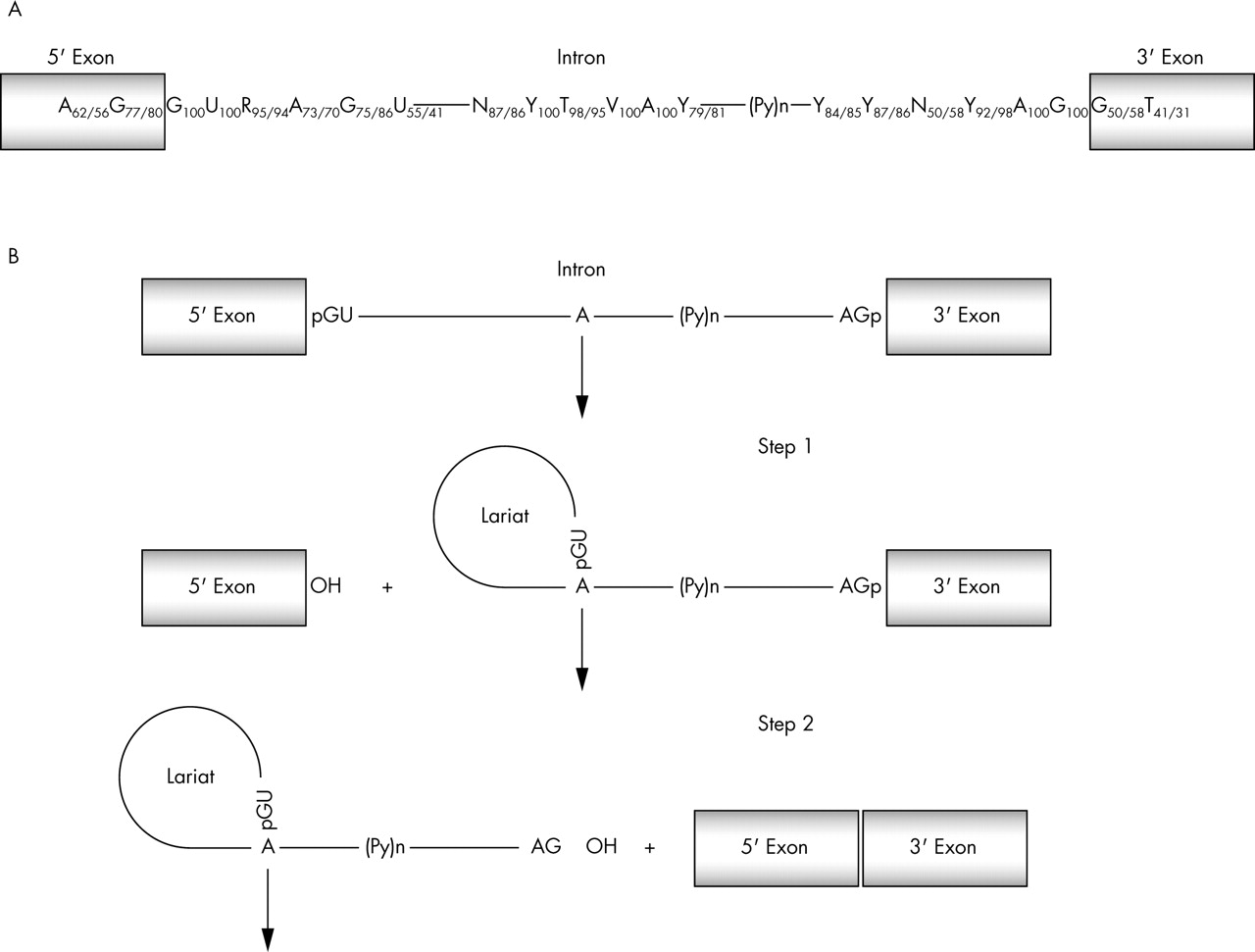
The remainder (0.73%) occurs in a lot of small groups (with a maximum size of 0.05%). Of these, 98.71% contain canonical GT-AG junctions (22 199 entries) and 0.56% have non-canonical GC-AG splice site pairs. 
The acceptor splice site is at the intron-exon boundary, which is expressed with consensus. After discarding sequences with putative errors and ambiguous location of splice junctions the verified dataset includes 22 489 entries. A splice site is a point where an exon and an intron intersect. Known EST sequences supported 22 815 of them. Sets of 38, 134, 81, and 96 native splice site contexts spanning a wide range of splicing ratios (Supplementary Fig. Splice site consensus sequences that drive exon recognition are located at the very termini of introns. We extracted 43 337 splice pairs from mammalian divisions of the gene-centered Infogene database, including sites from incomplete or alternatively spliced genes. A splice site mutation is a genetic mutation that inserts, deletes or changes a number of nucleotides in the specific site at which splicing takes place during the processing of precursor messenger RNA into mature messenger RNA. A database (SpliceDB) of known mammalian splice site sequences has been developed.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
